YaliFunTome
Yarrowia lipolytica Functional Screens of Tanscription Factor-ome Database
YALI number: YALI0B14443g
Number: TF009 | Associated name: JMC2 | NCBI: 2906736 | RefSeq DNA: XM_500884.1 | RefSeq Peptide: XP_500884.1 | UniProtKB: Q6CEM8
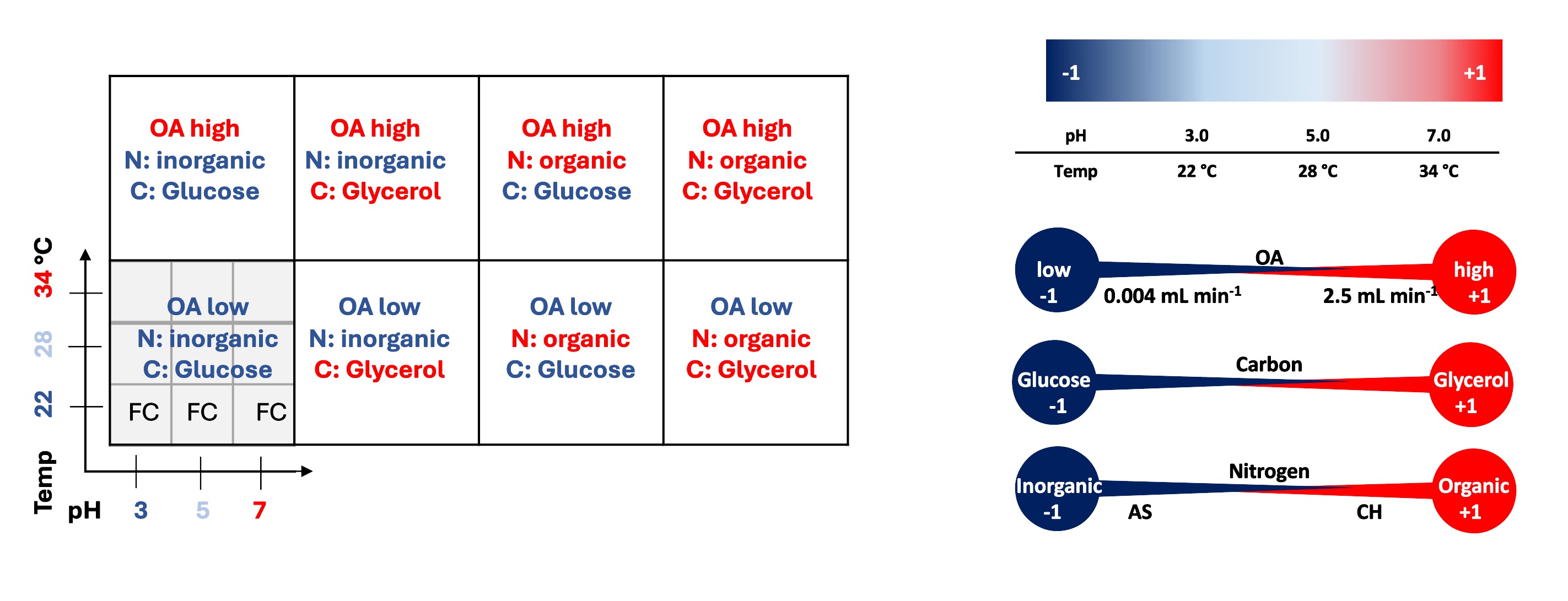
Fig.1. Graphical presentation of the arrangement of the variables' combinations corresponding to the data presentation scheme (left), and the variables' coding system (right). CH - casamino acid hydrolysate, AS - ammonium sulfate, OA- oxygen availability, FC - fold change of the response from the TF-OE strain over the control strain.
Legend:
FC - fold change over control strain, Sig - significant at p-value > 0.05, glu - glucose, gly - glycerol, i - inorganic, o - organic
| 2.0 | 1.7 | 1.5 | 1.3 | 1.1 | 1.0 | 0.7 | 0.5 | 0.3 | 0.1 |
Growth
Fold change in growth (optical density at 600 nm) of the TF-OE strain over the control strain
| 1.01 | ||
| 0.94 | 0.78 |
| 1.2 | 1.19 | |
| 1.29 |
| 0.92 | ||
| 2.07 | 0.95 |
| 1.04 | ||
| 1.28 | ||
| 0.9 |
| 1.08 | ||
| 1.69 | 1.06 | 1.16 |
| 1.8 | ||
| 1.36 |
Factor's Contribution
| YALI0B14443g |
|---|
| pH 7.14 |
| Temp 0.98 |
| Nitrogen 0.32↔ |
| OA -8.18 |
| Carbon non-significant |
Total r-Prot
Fold change in fluorescence of r-Prot in the TF-OE strain over the control strain
| 0.89 | ||
| 0.88 | 0.9 |
| 0.96 | 0.87 | |
| 1.05 |
| 0.72 | ||
| 1.11 | 0.9 |
| 0.9 | ||
| 0.94 | ||
| 0.82 |
| 1.22 | ||
| 1.15 | 0.89 | 0.99 |
| 1.69 | ||
| 1.09 |
Factor's Contribution
| YALI0B14443g |
|---|
| pH 5.74 |
| Temp -2.33 |
| OA -8.26 |
| Nitrogen non-significant |
| Carbon non-significant |
Normalized r-Prot
Fold change in fluorescence of r-Prot normalized per biomass in the TF-OE strain over the control strain
| 0.99 | 1 | |
| 0.71 | ||
| 0.68 | 0.97 | |
| 0.92 | 0.88 | |
| 0.97 |
| 0.82 | ||
| 0.88 | ||
| 0.73 | ||
| 0.84 | ||
| 0.72 | 1.12 | |
| 0.83 | ||
| 0.84 | 0.86 | |
| 0.95 | ||
Factor's Contribution
| YALI0B14443g |
|---|
| OA 2.45 |
| Temp -0.99 |
| Nitrogen -1.49 |
| pH -1.69 |
| Carbon non-significant |